May, 2022
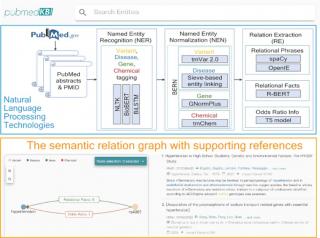
With the proliferation of genomic sequence data for biomedical research, the exploration of human genetic information by domain experts requires a comprehensive interrogation of large numbers of scientific publications in PubMed. However, a query in PubMed essentially provides search results sorted only by the date of publication. A search engine for retrieving and interpreting complex relations between biomedical concepts in scientific publications remains lacking. Here, we present pubmedKB, a web server designed to extract and visualize semantic relationships between four biomedical entity types: variants, genes, diseases, and chemicals. pubmedKB uses state-of-the-art natural language processing techniques to extract semantic relations from the large number of PubMed abstracts. Currently, over 2 million semantic relations between biomedical entity pairs are extracted from over 33 million PubMed abstracts in pubmedKB. pubmedKB has a user-friendly interface with an interactive semantic graph, enabling the user to easily query entities and explore entity relations. Supporting sentences with the highlighted snippets allow to easily navigate the publications. Combined with a new explorative approach to literature mining and an interactive interface for researchers, pubmedKB thus enables rapid, intelligent searching of the large biomedical literature to provide useful knowledge and insights. pubmedKB is available at https://www.pubmedkb.cc/.