Nov, 2015
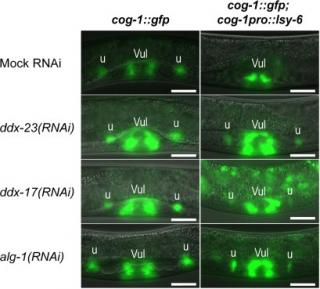
Primary microRNAs (pri-miRNAs) are cleaved by the nuclear RNase III Drosha to produce hairpin-shaped precursor miRNAs (pre-miRNAs). In humans, this process is known to be facilitated by the DEAD-box helicases p68 (DDX5) and p72 (DDX17). In this study, we performed a candidate-based RNAi screen in C. elegans to identify DEAD/H-box proteins involved in miRNA biogenesis. In a let-7(mg279) sensitized genetic background, knockdown of a homolog of yeast splicing factor Prp28p, DDX-23, or a homolog of human helicases p68 and p72, DDX-17, enhanced let-7 loss-of-function phenotypes, suggesting that these helicases play a role in let-7 processing and/or function. In both ddx-23(RNAi) and ddx-17(RNAi), levels of mature let-7 were decreased while pri-let-7 was found to accumulate, indicating that the helicases likely act at the level of pri-let-7 processing. DDX-23 and DDX-17 were also required for the biogenesis of other known heterochronic miRNAs, including lin-4 and the let-7 family members miR-48, miR-84 and miR-241. Their function was not confined to the heterochronic pathway, however, since they were both necessary for down-regulation of cog-1 by the spatial patterning miRNA, lsy-6. Here, we present a novel function for C. elegans DDX-23 in pri-miRNA processing, and also suggest a conserved role for DDX-17 in this process.